 These methods are well established and result in high quality DNA or RNA, but they are often very laborious, time-consuming and cost-intensive, or suffer from insufficient and inconsistent yields of NAs. Hazardous chemicals such as phenol and chloroform 10, or commercial kits are then used for NA purification, depending on the area of application and the matrix in which the cells are investigated. State-of-the-art extraction procedures for microorganisms traditionally use incubation steps with enzymes such as lysozyme and proteinase K to digest cell wall components and interfering proteins, respectively. The purification of the NAs is, in most cases, achieved either by precipitation followed by washing steps, or by column-based purification protocols. Common methods for cell lysis involve thermal, chemical, enzymatic, or mechanical treatment of the cells or a combination of those 1. These steps typically involve the lysis of the cells, the purification of the NAs to remove other cell components, inhibitory substances or degrading enzymes, and the subsequent recovery of the desired NAs. 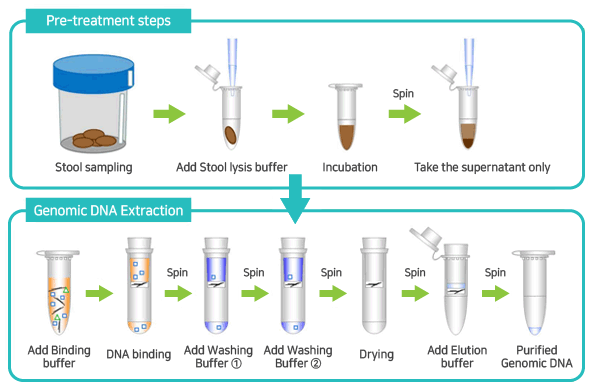
However, to detect the desired sequence of a certain NA (DNA or RNA), preceding steps are necessary to isolate the genetic material from the respective cells. The field of microbial molecular diagnostics comprises various methods for the specific detection of nucleic acids (NAs) from different microorganisms 1, 2 – e.g., human pathogens in clinical and environmental samples 3, 4, 5, faecal indicator bacteria in water 6, 7, or harmful microbial agents in food and feed 8, 9. This makes the method ideal for high sample throughput and offers the opportunity for DNA extraction from bacteria in resource-limited settings or even in the field. The final protocol for DNA extraction using the two ILs is very low-cost, avoids the use of hazardous chemicals and can be performed in five minutes on a simple heating block. In comparison to the reference method, extraction yields of the IL-based procedure were within one order of magnitude for most of the strains. 
All methods were evaluated on four Gram-positive and four Gram-negative bacterial species that are highly relevant for environmental, food, or clinical diagnostics. 
The two best ILs, choline hexanoate and 1-ethyl-3-methylimidazolium acetate, were compared with the reference enzymatic method and two commercial DNA extraction kits. The cell lysis potential of different IL/buffer combinations was assessed by application on Enterococcus faecalis as a model organism for Gram-positive bacteria. First, we tested eight ILs in different buffer systems for their inhibitory effects on quantitative PCR. We developed a method for the rapid preparation of genomic DNA from bacteria based on hydrophilic ionic liquids (ILs). The extraction of nucleic acids from microorganisms for subsequent molecular diagnostic applications is still a tedious and time-consuming procedure. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
